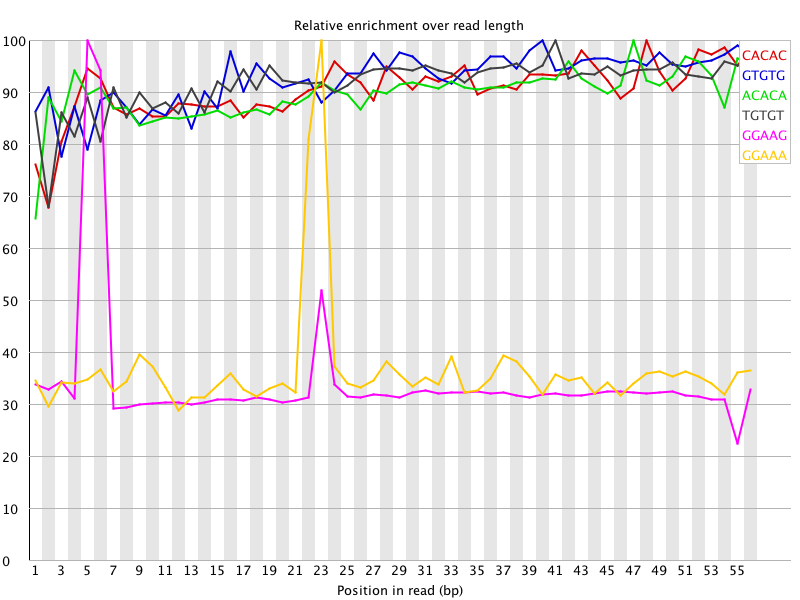
I ran FatsQC on a set of 4 Illumina PE exomes and got really weird graphs for the kmer distribution. I've never seen this pattern before - does anyone know what is going on and what to do about it?
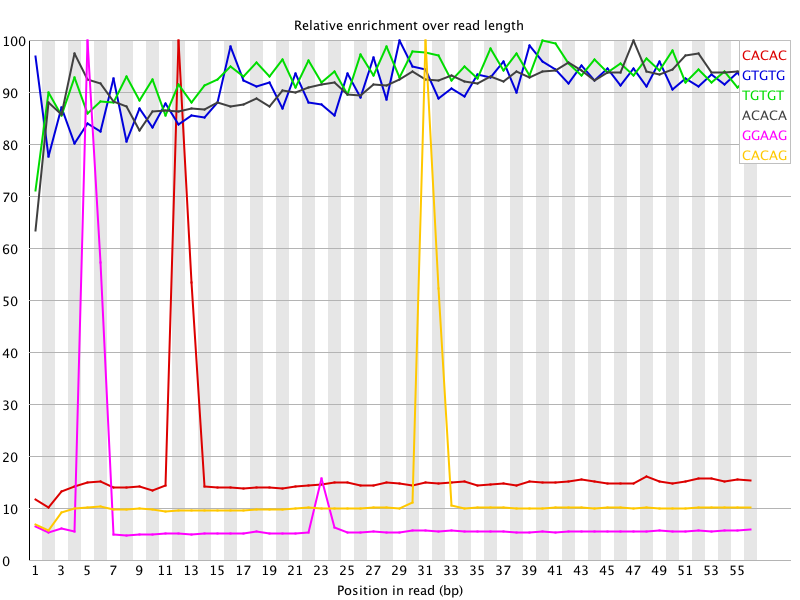
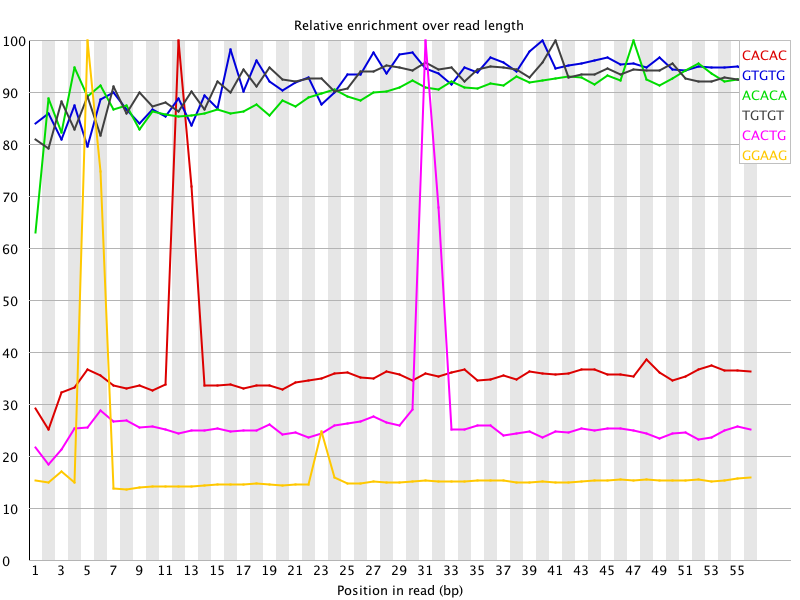
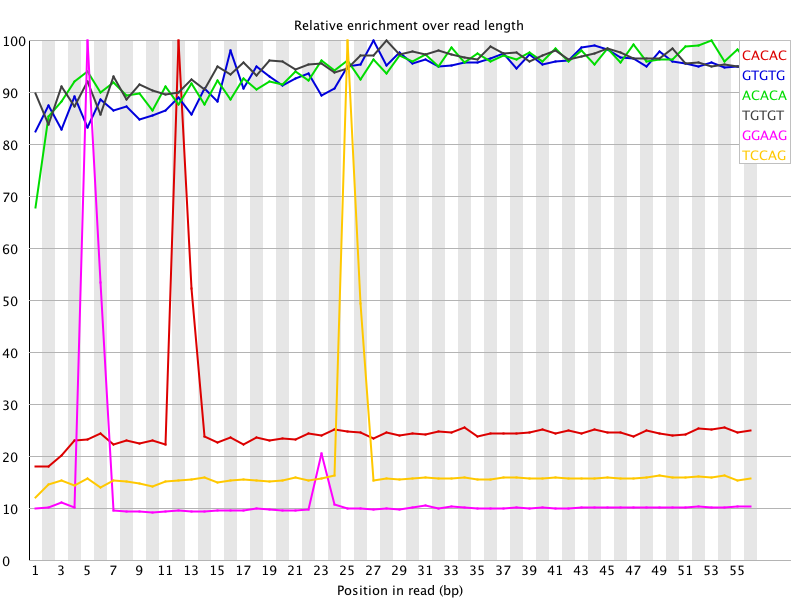
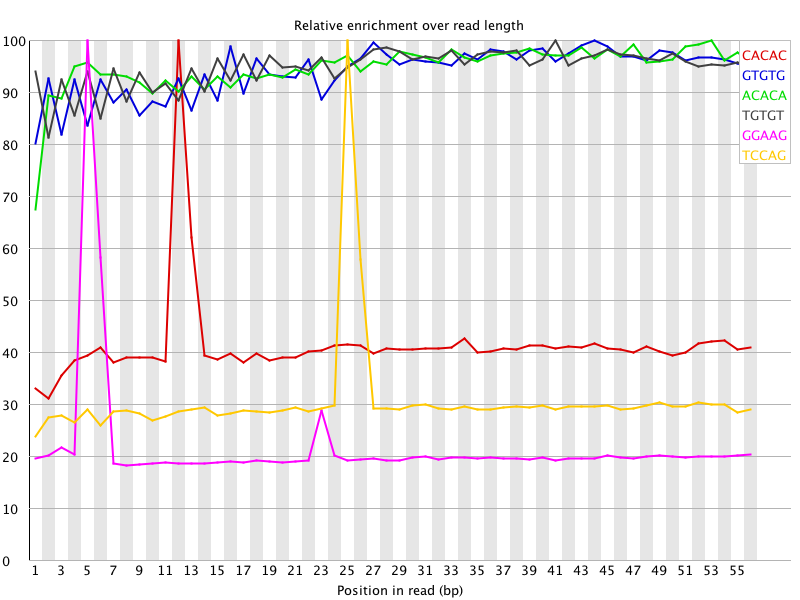




You are currently viewing the SEQanswers forums as a guest, which limits your access. Click here to register now, and join the discussion
| Topics | Statistics | Last Post | ||
|---|---|---|---|---|
|
Started by seqadmin, Today, 08:47 AM
|
0 responses
9 views
0 likes
|
Last Post
by seqadmin
Today, 08:47 AM
|
||
|
Started by seqadmin, 04-11-2024, 12:08 PM
|
0 responses
60 views
0 likes
|
Last Post
by seqadmin
04-11-2024, 12:08 PM
|
||
|
Started by seqadmin, 04-10-2024, 10:19 PM
|
0 responses
57 views
0 likes
|
Last Post
by seqadmin
04-10-2024, 10:19 PM
|
||
|
Started by seqadmin, 04-10-2024, 09:21 AM
|
0 responses
53 views
0 likes
|
Last Post
by seqadmin
04-10-2024, 09:21 AM
|
Comment