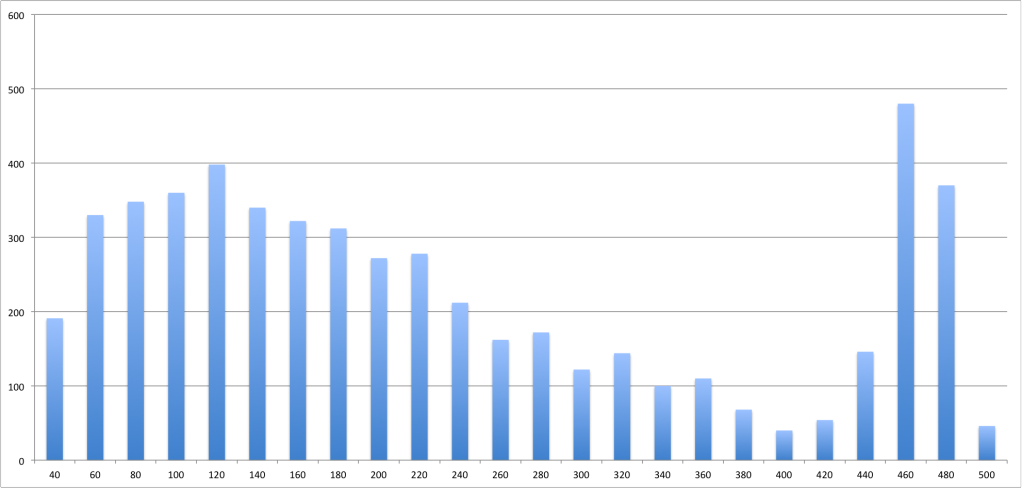
I see in many bioinformatics papers assumptions that the fragment lengths for Illumina data are be normally distributed. I've seen some datasets for which this doesn't seem to be the case, however. I've seen what look like very skewed and bimodal distributions in some of the 1000 Genomes Project data.
I'm a computer scientist, so I don't know much about what to expect this data to look like or why it would have a given fragment length distribution. I've been searching for the past couple days for a reference, but I've come up empty.
Is there anyone here that can help me understand this better or point me to a resource where I could learn more? Any help would be greatly appreciated!
I'm a computer scientist, so I don't know much about what to expect this data to look like or why it would have a given fragment length distribution. I've been searching for the past couple days for a reference, but I've come up empty.
Is there anyone here that can help me understand this better or point me to a resource where I could learn more? Any help would be greatly appreciated!
Comment