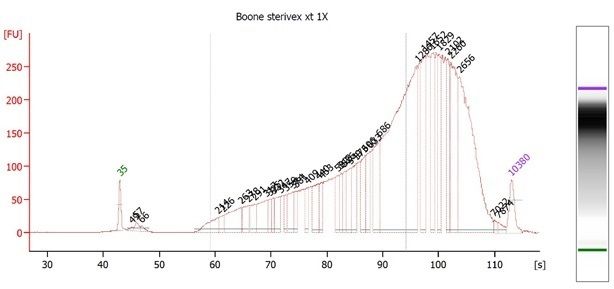
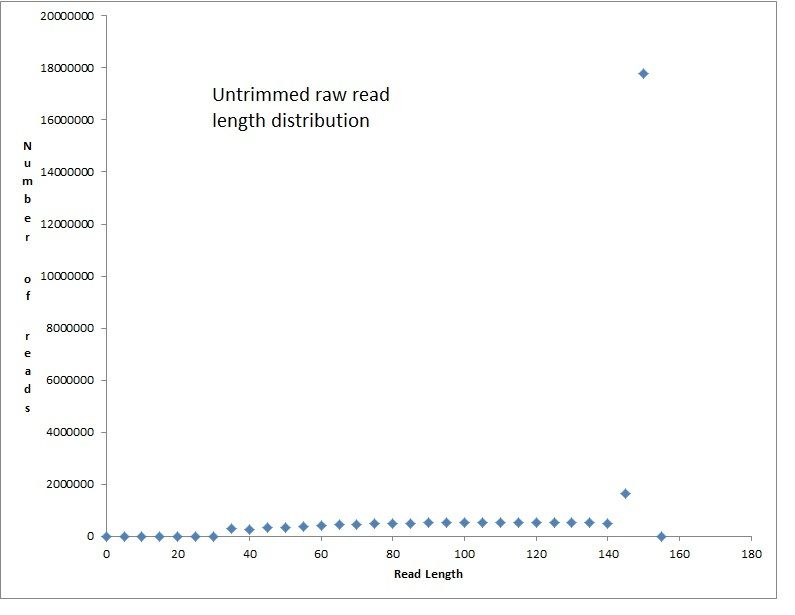
Hi all - I am very new to metagenomics analyses, so please bear with me. I have a metagenome generated from 2L of Sterivex filtered stream water run on a NextSeq500 using NexteraXT library prep, which was dual indexed and run alongside other samples. The number of raw reads was 29,570,310 paired end reads (150bpX2), which I know is not necessarily a lot for a metagenome. In the following code snippets below I did not use the real file names since I have 4 sets of merged reads, and four sets of unmerged reads (2 files for each of four lanes) with long names that I need to shorten, but the commands worked with the real filenames. For read processing I used bbduk and bbmerge as follows:
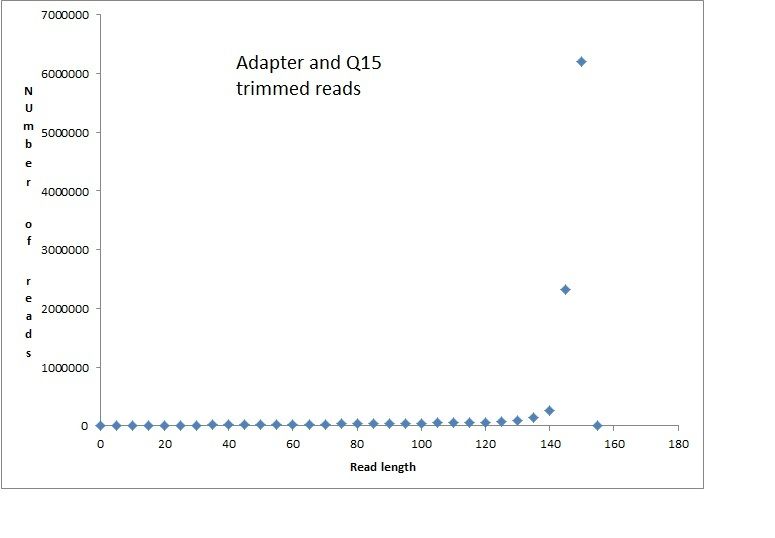
Adapter and Quality trimming:
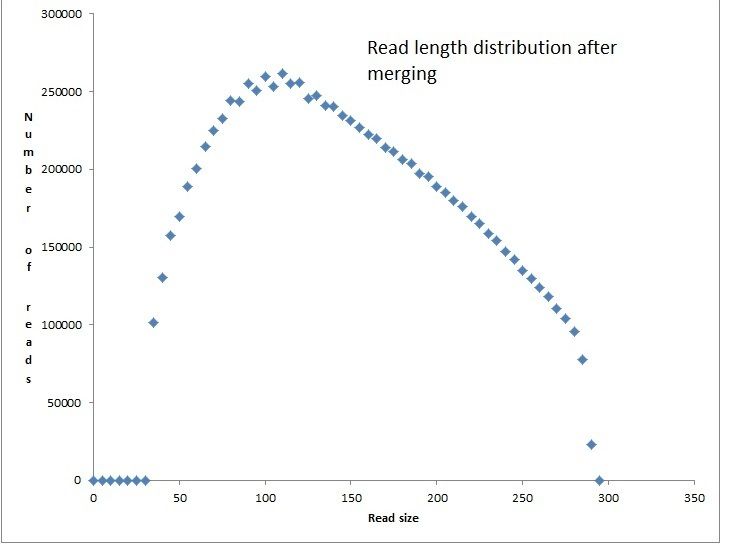
Merging overlapping pairs:
I have taken these quality trimmed reads and submitted both the merged and unmerged read pairs to MG-RAST. I also want to do a de novo assembly, which is largely where my questions arise. I used Megahit as follows:
I ended up with:
- 40,747 contigs totalling 19663614 bp
- min contig size of 200, max size of 7816 bp, avg 483 bp
- N50 = 458 bp
From what I have read, that N50 is low and likely reflects insufficient coverage of the sampled community. I should be shooting for N50 >= 1000 correct? At this point, additional sequencing is not an option, so I am trying to work with what I have.
Now that I have contigs from a de novo assembly, I should have a large proportion of reads not incorporated into those contigs that are still valuable potentially for taxonomic and functional annotation. From the megahit assembly, I do not get a subset of reads that were not incorporated into the contigs. Therefore I followed the tutorial at the following url for getting unassembled reads:
I mapped the reads as follows:
Then, I output coverage per contig using pileup.sh from the bbmap suite as follows:
The following is the first few lines of the coverage file:
Taking the first contig k127_4 as an example, the coverage (Covered_percent) listed is 0. However, at least some reads had to be joined to make that contig. Shouldn't each contig have a coverage greater than zero? I have output a list of unmapped reads as follows (admittedly at this point I am just following the Megahit tutorial, and am still learning what the options mean for Samtools):
How can I be sure these unmapped reads are not part of the assembled contigs when the coverage listed for many of the contigs is zero? My end goal here is a list of contigs, and a list of unmapped reads that can both be Blastx'd using DIAMOND and visualized in MEGAN. If this is the wrong approach to attain this goal, I would appreciate any guidance or comments. Thanks for your time -
Adapter and Quality trimming:
Code:
>bbduk.sh in1=read1.fq in2=read2.fq out1=name_R1_Q15_k29.fq out2=name_R2_Q15_k29.fq ref=/usr/local/bbmap/resources/nextera.fa qtrim=r trimq=15 k=29 mink=11 hdist=1 ktrim=r tpe tbo
Code:
>bbmerge.sh in1=name_R1_Q15_k29.fq in2=name_R2_Q15_k29.fq out=name_Q15_K29_merged.fq outu=name_Q15_k29_unmerged.fq mininsert=20
Code:
>./megahit --12 name_Q15_k29_unmerged.fq -r name_Q15_k29_merged.fq --k-max=127 -o output_directory
- 40,747 contigs totalling 19663614 bp
- min contig size of 200, max size of 7816 bp, avg 483 bp
- N50 = 458 bp
From what I have read, that N50 is low and likely reflects insufficient coverage of the sampled community. I should be shooting for N50 >= 1000 correct? At this point, additional sequencing is not an option, so I am trying to work with what I have.
Now that I have contigs from a de novo assembly, I should have a large proportion of reads not incorporated into those contigs that are still valuable potentially for taxonomic and functional annotation. From the megahit assembly, I do not get a subset of reads that were not incorporated into the contigs. Therefore I followed the tutorial at the following url for getting unassembled reads:
I mapped the reads as follows:
Code:
>bbwrap.sh ref=name_sterivex_assembly/final.contigs.fa in=name_Q15_k29_unmerged.fq,name_Q15_k29_merged.fq out=aln.sam.gz kfilter=22 subfilter=15 maxindel=80
Code:
>pileup.sh in=aln.sam.gz out=cov.txt
The following is the first few lines of the coverage file:
Code:
aaron@aaron-VirtualBox:~/PineCreek/Boone_sterivex_2014-30975030$ head cov.txt #ID Avg_fold Length Ref_GC Covered_percent Covered_bases Plus_reads Minus_reads Read_GC k127_4 flag=1 multi=2.0000 len=391 0.0000 391 0.0000 0.0000 0 0 0 0.0000 k127_5 flag=1 multi=2.0000 len=414 0.0000 414 0.0000 0.0000 0 0 0 0.0000 k127_6 flag=1 multi=1.0000 len=440 0.3432 440 0.0000 34.3182 151 0 1 0.6424
Code:
>samtools view -u -f4 aln.sam.gz | samtools bam2fq -s unmapped.se.fq - > unmapped.pe.fq

Comment