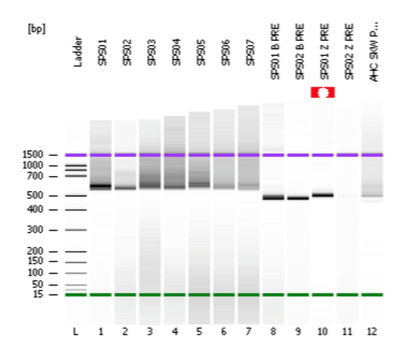
I've been making two-step amplicon libraries for years now with very few problems. The original amplicons are tailed with partial Nextera xt adapters and then I use Phusion HF to add the rest of the adapter with 6 cycles of PCR and KAPA Pure beads for cleanup. Recently, the yield has been much lower than expected and, rather than a pretty clean peak on the Bioanalyzer, there's been a lot of high molecular weight smearing. The samples labeled SPS01-07 are after the second round of amplication while the samples labeled Pre are before. Suggestions as to what might be going on?
Seqanswers Leaderboard Ad
Collapse
Announcement
Collapse
No announcement yet.
X
-
Hey there, microgirl:
I was wondering if you tried what luc suggested and if this worked.
Also, do you have a low annealing temperature on your PCR cycles? I figure it's stayed the same, but since other things have been changed, maybe the thermocycler programming needs to be altered as well.
I've only been doing library prep for around 1.5 yr, but it surprised me how low the annealing temps in a lot of protocols are set to. We anneal at 50C!
Comment
Latest Articles
Collapse
-
by seqadmin
The field of epigenetics has traditionally concentrated more on DNA and how changes like methylation and phosphorylation of histones impact gene expression and regulation. However, our increased understanding of RNA modifications and their importance in cellular processes has led to a rise in epitranscriptomics research. “Epitranscriptomics brings together the concepts of epigenetics and gene expression,” explained Adrien Leger, PhD, Principal Research Scientist...-
Channel: Articles
04-22-2024, 07:01 AM -
-
by seqadmin
Proteins are often described as the workhorses of the cell, and identifying their sequences is key to understanding their role in biological processes and disease. Currently, the most common technique used to determine protein sequences is mass spectrometry. While still a valuable tool, mass spectrometry faces several limitations and requires a highly experienced scientist familiar with the equipment to operate it. Additionally, other proteomic methods, like affinity assays, are constrained...-
Channel: Articles
04-04-2024, 04:25 PM -
ad_right_rmr
Collapse
News
Collapse
| Topics | Statistics | Last Post | ||
|---|---|---|---|---|
|
Started by seqadmin, Today, 08:47 AM
|
0 responses
11 views
0 likes
|
Last Post
by seqadmin
Today, 08:47 AM
|
||
|
Started by seqadmin, 04-11-2024, 12:08 PM
|
0 responses
60 views
0 likes
|
Last Post
by seqadmin
04-11-2024, 12:08 PM
|
||
|
Started by seqadmin, 04-10-2024, 10:19 PM
|
0 responses
59 views
0 likes
|
Last Post
by seqadmin
04-10-2024, 10:19 PM
|
||
|
Started by seqadmin, 04-10-2024, 09:21 AM
|
0 responses
54 views
0 likes
|
Last Post
by seqadmin
04-10-2024, 09:21 AM
|
Comment