Hello,
I have a question: For heterozygous deletion, Sanger sequencing chromatograms will look like what?
This is because, I found heterozygous deletion in 1 sample when i did ion torrent sequencing. Brief information about that mutation is below with IGV figure.
Chr Start End Ref Alt Func.refGene Gene.refGene
chr1 155208272 155208276 AGGGC AGGC intronic GBA
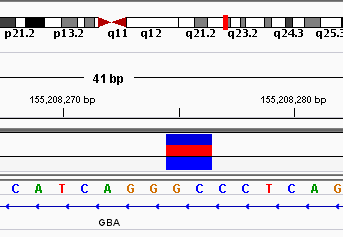
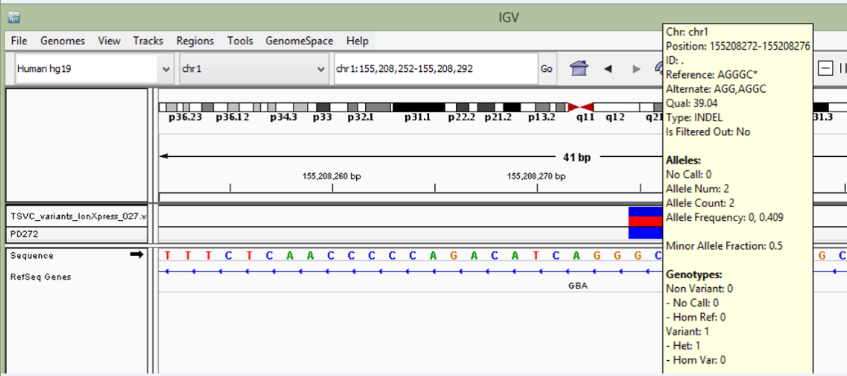

I validate this mutation using Sanger sequencing and I got this result. Does this result correct? Does it is represent heterozygous deletion?
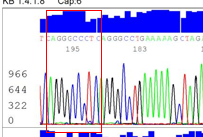
I have a question: For heterozygous deletion, Sanger sequencing chromatograms will look like what?
This is because, I found heterozygous deletion in 1 sample when i did ion torrent sequencing. Brief information about that mutation is below with IGV figure.
Chr Start End Ref Alt Func.refGene Gene.refGene
chr1 155208272 155208276 AGGGC AGGC intronic GBA
I validate this mutation using Sanger sequencing and I got this result. Does this result correct? Does it is represent heterozygous deletion?
Comment