Hi,
I was using FASTQC to QC my directional mRNA-Seq data obtained from suspension culture cells. I have about 30 million reads.
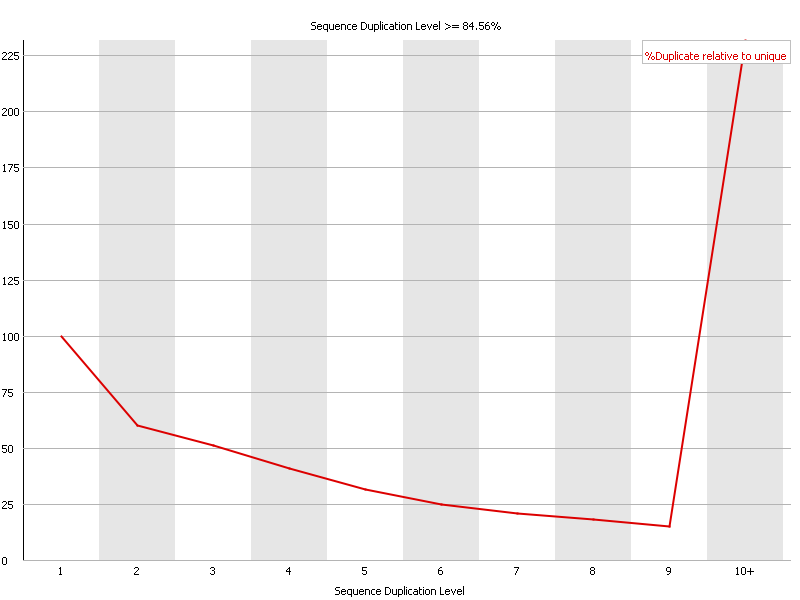
Although most of my QC stats are fine, I see a big uptick in the "Duplicate sequences" section of sequences with duplication levels > 10 (see below). Sequence Duplication Level >84.56%.
I was wondering what could be wrong. There were 2 possibilities I could think of:
1) Some amplification bias in PCR and/or
2) Since the RNA is not very diverse (its from suspension cells - same cell type) and sequenced to a high coverage, many sequences got sequenced multiple times.
Wonder if the second reason makes sense? If it is true, by extension, it also means that we have successfully sequenced even the very low abundance transcripts. However, if it was PCR bias, that wouldnt be true. Wonder if there is a way to distinguish between these two possibilities?
I'd appreciate any suggestions.
Thanks
I was using FASTQC to QC my directional mRNA-Seq data obtained from suspension culture cells. I have about 30 million reads.
Although most of my QC stats are fine, I see a big uptick in the "Duplicate sequences" section of sequences with duplication levels > 10 (see below). Sequence Duplication Level >84.56%.
I was wondering what could be wrong. There were 2 possibilities I could think of:
1) Some amplification bias in PCR and/or
2) Since the RNA is not very diverse (its from suspension cells - same cell type) and sequenced to a high coverage, many sequences got sequenced multiple times.
Wonder if the second reason makes sense? If it is true, by extension, it also means that we have successfully sequenced even the very low abundance transcripts. However, if it was PCR bias, that wouldnt be true. Wonder if there is a way to distinguish between these two possibilities?
I'd appreciate any suggestions.
Thanks

Comment