Hi all,
Really hope for your help, guys.
I finally got my ChIP-Seq data back. Hoping it is not too bad. You can check for the history of my libraries prep here:
Now, I want to analyse the data by myself as much as I can. Unfortunately due to the cost, attending a workshop is out of question.. An online course, possibly. If you know one, please share with me.
The question is what would be the best approach to analyse my data? What do I need?
I know R and I was participating in assembling a genome "on cloud", but I do not remember much. For the best of my memory I will not be able to work with my data on my computer. I have 20 files (forward and reverse reads) about 3.5 Gb zipped.
I tried to load them on galaxy via FTP, but it is painfully slow (~24h each! file). Then, some files possibly got corrupted - FastQC in galaxy fails on them.
I found a couple of resources to read:
a list on aps on biostars:
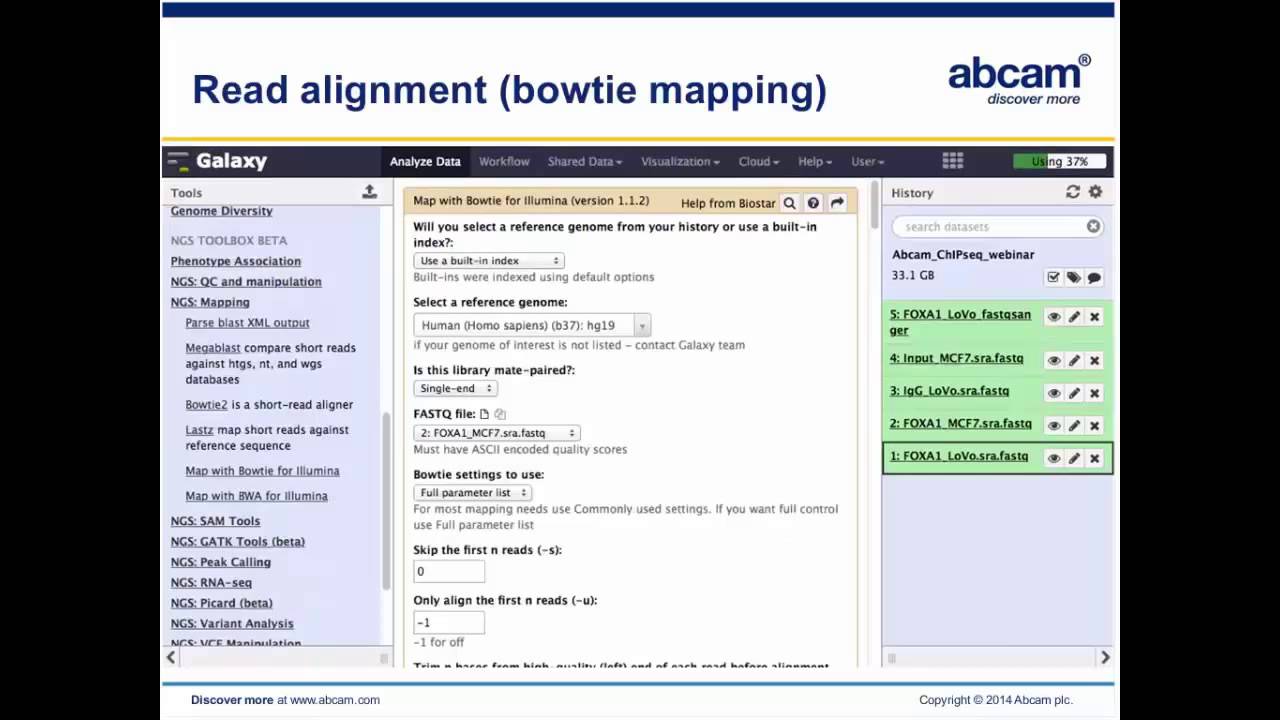
Using Galaxy. A workshop from Abcam:
and
ChIP data with Galaxy. From Galaxy:
and
Using R:
It looks like uploading to galaxy will never finish. Does it make sense to try to do everything in R? What is the most common pipeline used for NGS data analysis.
If I will not be able to work with the files on my computer, does anybody know how to organise that "cloud commuting" in detail? I have Amazon cloud account and virtual box. What else and how?
Thank you!
Really hope for your help, guys.
I finally got my ChIP-Seq data back. Hoping it is not too bad. You can check for the history of my libraries prep here:
Now, I want to analyse the data by myself as much as I can. Unfortunately due to the cost, attending a workshop is out of question.. An online course, possibly. If you know one, please share with me.
The question is what would be the best approach to analyse my data? What do I need?
I know R and I was participating in assembling a genome "on cloud", but I do not remember much. For the best of my memory I will not be able to work with my data on my computer. I have 20 files (forward and reverse reads) about 3.5 Gb zipped.
I tried to load them on galaxy via FTP, but it is painfully slow (~24h each! file). Then, some files possibly got corrupted - FastQC in galaxy fails on them.
I found a couple of resources to read:
a list on aps on biostars:
Using Galaxy. A workshop from Abcam:
and
ChIP data with Galaxy. From Galaxy:
and
Using R:
It looks like uploading to galaxy will never finish. Does it make sense to try to do everything in R? What is the most common pipeline used for NGS data analysis.
If I will not be able to work with the files on my computer, does anybody know how to organise that "cloud commuting" in detail? I have Amazon cloud account and virtual box. What else and how?
Thank you!



 ) wifi is much slower. In particular from a private (home) network.
) wifi is much slower. In particular from a private (home) network.
Comment