Hi,
It is my first time to use the house-make reagents to prepare DGE-tag profiling library for sequencing. The sequencing result seem to not perfectly comfirm the microarray result and they are from the same samples (see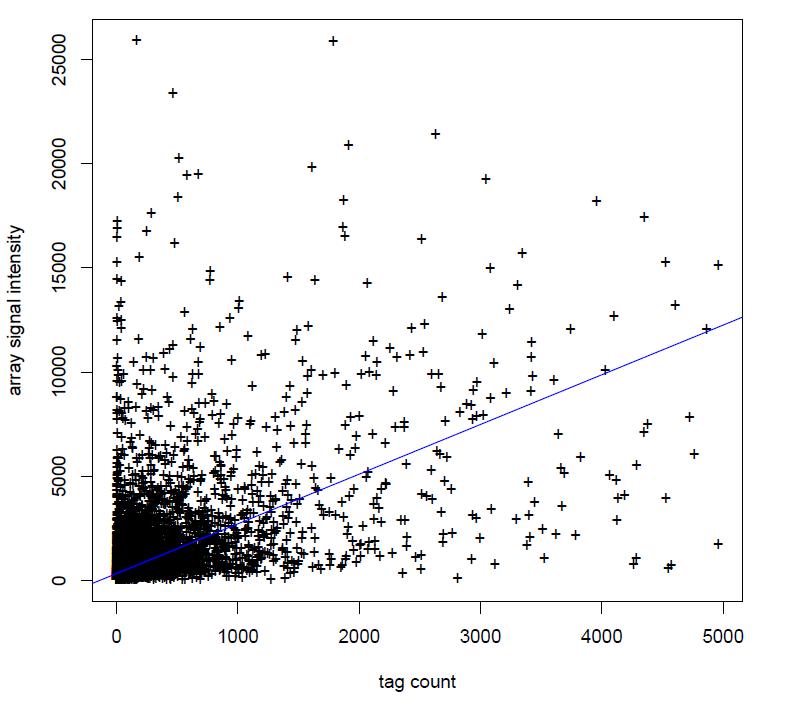 ). The eighty percent reads can map to the arabidopsis cDNA sequences, but only about fifty percent reads can map to tag table from the nearest restriction site to 3' teminal of cDNA. The tag table made by myself. Is it normal? Could you give me some advice on sample preparation?
). The eighty percent reads can map to the arabidopsis cDNA sequences, but only about fifty percent reads can map to tag table from the nearest restriction site to 3' teminal of cDNA. The tag table made by myself. Is it normal? Could you give me some advice on sample preparation?
Maybe the insufficiency of enzyme digestion result in the low proportion of reads mapping to tag table. But I don't think it's the main reason of the low consistency with microarray data.
Thanks.
It is my first time to use the house-make reagents to prepare DGE-tag profiling library for sequencing. The sequencing result seem to not perfectly comfirm the microarray result and they are from the same samples (see
Maybe the insufficiency of enzyme digestion result in the low proportion of reads mapping to tag table. But I don't think it's the main reason of the low consistency with microarray data.
Thanks.

Comment